BUGS has a “zeros trick” (Lund, Jackson, Best, Thomas, Spiegelhalter, 2013, pages 204-206; see also an online illustration) for specifying a new distribution which is not in the standard set. The idea is to couple an invented-for-the-moment Poisson density to observations of structural zeros and transform the component sought in the overall likelihood (for the MCMC) so it comes out to be both positive and the form wanted. The steps are, essentially:
z[i]<- 0 z[i] ~ dpois(phi[i]) phi[i]<- -log(L[i]) + constant L[i]<- ...
borrowing the code from Lund, et al. Here L[i] is the likelihood sought, and “constant” is the constant included to be sure phi[i] remains positive. Then, paraphrasing Lund, et al again, L[i] is set to a function of y[i] and proportional to the likelihood
.
Now in JAGS you can’t write
z[i]<- 0 z[i] ~ dpois(phi[i])
because you appear to be redefining z[i]. Instead z[i] can be put in a “data section” like
data
{
for (i in 1:n)
{
z[i]<- 0
}
}
and then in the model section
model
{
for (i in 1:n)
{
...
z[i] ~ dpois(phi[i])
...
}
}
But there are other neat things this construct let’s you do.
I was working on implementing “error in variables” problems in JAGS, inspired in part by Congdon’s Section 15.2 in his Bayesian Statistical Modelling, 2nd edition (2006), in part by blog posts by Maxwell Joseph of Ecology in Silico here and here, and in part by a question from a colleague. I had studied so-called “total least squares” using an SVD in Golub and Van Loan’s Matrix Computations (in a course I took taught by Van Loan himself), but I wanted to do this using a Bayesian approach. I found it hard to set up to do using other than a home-grown MCMC sampler, and I’ve learned to respect people who build such samplers well and thought I shouldn’t repeat that. Still, I found no easy way to do this in general in R‘s MCMCPack and the like. I decided I needed to learn BUGS. I did.
A reparameterization of solving a simple affine linear model with errors in covariate and response led to an expression for residuals like
and I very much wanted to write something like
xobs[j,i] * sin(angle) - yobs[j,i] * cos(angle) + b * cos(angle) ~ dnorm(0, 10000)
But this is illegal in JAGS, since left hand sides cannot be such complicated expressions. Unlike WinBUGS, you can’t even write “log(x) ~ …”. Zeros trick to the rescue! With things like z[i] above, say, r[j,i], even if the expression sought is not permitted, you can write
r[j,i] ~ dnorm(xobs[j,i] * sin(angle) - yobs[j,i] * cos(angle) + b * cos(angle), 10000)
Recall that in JAGS and BUGS, dnorm(a,b) is a density with a mean a and a precision b, not a variance. Precision is reciprocal variance, so a precision of 10000 is a standard deviation of 0.01, or small. As in the case of the zeros trick for defining another likelihood, this constrains r[j,i] to be nearly exactly the value of the mean, and since there’s no “room” for variation in r[j,i], and since xobs[j,i] and yobs[j,i] are also given as data, this constraint “backs up” into constraints on “angle” and “b”, defined with priors separately. Note there’s a specific relationship here defined among these quantities.
But this does not need to stop with error-in-variables problems. Any time the zero-crossings or roots of an expression are sought, it can be set up to be found stochastically by using the same expression. In short, MCMC is being used to search for the roots and, with general enough priors, like a Jeffrey’s prior, and enough of a run, the multivariate posterior that will be returned will contain evidence for all the roots found.
I find that very cool. It’s not at all an original idea, and James Spall talks about it in his book Introduction to Stochastic Search and Optimization (Wiley, 2003).
Incidently, as in most computational situations, I found JAGS quite fast when solving a number of the problems in Congdon’s Section 15.2, and also the problem at Ecology in Silico. I did not formulate the model the same way, however, and so it’s hard to say exactly what the difference is between the JAGS version Maxwell Joseph tried, the STAN version he reported efficient execution with, and this one. I’m including the entire JAGS code for the case below, just for reference. Note the code needs the value of and I pass this in from R. I ran this using:
dataList<- list(n=n, nobs=nobs, xobs=t(xobs), yobs=t(yobs), Pi=pi)
print(dataList)
#------------------------------------------------------------------------------
# INTIALIZE THE CHAINS.
initialization<- function(chain)
{
return(list(.RNG.name="base::Mersenne-Twister",
angle.theta=rbeta(1, 0.5, 0.5),
b.theta=rbeta(1, 0.5, 0.5),
sigma.x=rgamma(1, shape=3, rate=0.5),
sigma.y=rgamma(1, shape=3, rate=0.5),
x.theta=rbeta(n, 0.5, 0.5),
y.theta=rbeta(n, 0.5, 0.5)))
}
# Run the model and produce plots
results<- run.jags(model=modelstring,
silent.jags=FALSE,
datalist=dataList,
n.chains=10,
inits=NA,
initlist=initialization,
burnin=3000, sample=5000,
adapt=1000, thin=3,
summarise=TRUE, confidence=0.98,
plots=TRUE, psrf.target=1.05,
normalise.mcmc=TRUE, check.stochastic=TRUE,
check.conv=TRUE, keep.jags.files=TRUE,
jags.refresh=0.1, batch.jags=FALSE,
method="parallel")
var Pi, n, nobs, yobs[nobs, n], xobs[nobs, n], y[n],
y.theta[n], b, b.theta, tan.angle, angle.theta,
angle, x[n], x.theta[n], sigma.x, sigma.y, r[nobs, n];
data
{
for (i in 1:n)
{
for (j in 1:nobs)
{
r[j,i]<- 0
}
}
}
model
{
tan.angle<- tan(angle)
# priors
b.theta ~ dbeta(0.5, 0.5) # Jeffrey's prior
b<- logit(b.theta) # intercept
angle.theta ~ dbeta(2,2)
angle<- Pi*2*(angle.theta - 0.5)/2 # angle w.r.t. abscissa
# precisions
sigma.x ~ dgamma(3, 0.5)
sigma.y ~ dgamma(3, 0.5)
tau.x<- pow(sigma.x, -2)
tau.y<- pow(sigma.y, -2)
for (i in 1:n)
{
x.theta[i] ~ dbeta(0.5, 0.5) # Jeffrey's prior
x[i]<- logit(x.theta[i])
y.theta[i] ~ dbeta(0.5, 0.5) # Jeffrey's prior
y[i]<- logit(y.theta[i])
}
# model structure
for (i in 1:n)
{
for (j in 1:nobs)
{
r[j,i] ~ dnorm(xobs[j,i] * sin(angle) - yobs[j,i] * cos(angle) + b * cos(angle), 10000)
xobs[j, i] ~ dnorm(x[i], tau.x)
yobs[j, i] ~ dnorm(y[i], tau.y)
}
}
#inits# .RNG.name, angle.theta, b.theta, x.theta, y.theta
#monitor# sigma.y, sigma.x, tan.angle, b
#data# n, nobs, xobs, yobs, Pi
}






Pingback: Less evidence for a global warming hiatus | Hypergeometric
Dears
I have question on Zeros tricks coding of generalized error distribution with mean a precision b and shape parameter p.
How can write zeros trick for GED(a,b,p)
JAGS 4 (and predecessors) does not have an implementation of the Generalized Normal Distribution (= GED). Your first challenge will be simulating that with the built-in Standard Normal and the built-in Double Exponential. Assuming that’s done, and assuming is your shape parameter, you’ll need to define a hyperprior for
is your shape parameter, you’ll need to define a hyperprior for  in addition to hyperpriors for
in addition to hyperpriors for  and
and 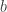 .
.
A way of simulating the Generalized Normal, assuming you are interested only in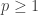 , is to treat it as a mixture of a Laplace distribution and a Uniform. The Laplace in JAGS is called the Double Exponential distribution (see the JAGS user manual). Then, if
, is to treat it as a mixture of a Laplace distribution and a Uniform. The Laplace in JAGS is called the Double Exponential distribution (see the JAGS user manual). Then, if  is a mixing parameter, having, perhaps, a Beta hyperprior, the resulting r.v. would be
is a mixing parameter, having, perhaps, a Beta hyperprior, the resulting r.v. would be  , where
, where 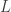 denotes your simulated Double Exponential and
denotes your simulated Double Exponential and 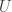 your simulated Uniform. Alternatively, you could do something ad hoc and use
your simulated Uniform. Alternatively, you could do something ad hoc and use 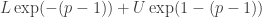 perhaps using a Gamma hyperprior on
perhaps using a Gamma hyperprior on  . I would expect mixing and convergence to be slow. See the JAGS user manual for suggestions on how to rewrite expressions so to center them and improve convergence. Also, choosing bounds on the Uniform will need to be guided by your application, as will sensible ranges for the hyperpriors.
. I would expect mixing and convergence to be slow. See the JAGS user manual for suggestions on how to rewrite expressions so to center them and improve convergence. Also, choosing bounds on the Uniform will need to be guided by your application, as will sensible ranges for the hyperpriors.
Alternatively, chuck the Generalized Error Distribution altogether, and try to use a mixture of Normal distributions and the JAGS builtin-in dnormmix and its mix module. (See the JAGS manual for version 4, section 4.3.) Whether you can get away doing this or not will depend upon what, specifically, your problem is. Scaling and ranges of r.v. matter.
Pingback: Blind Bayesian recovery of components of residential solid waste tonnage from totals data | Hypergeometric